MAE 127: Lecture 6
More on Matlab
The goals for this computer session are to help you learn some
additional Matlab tricks that will help you out with real data
analysis, including providing guidance for the problems that you
encounter in this class.
Data formats. Loading other
data: Last time we loaded Matlab binary files (with
the
suffix .mat) using:
load
sample_data.mat
load('sample_data.mat')
But not all data is in Matlab binary format.
For plain text (or ASCII) files, Matlab also loads data fairly
automatically:
load
sample_data.ascii
DATA=load('sample_data.ascii'); and
DATA2=load('lecture3.timeseries');
DATA=load('-ascii','sample_data.ascii');
What about Microsoft Word files or Excel spreadsheets? These have
binary data formating that Matlab is not designed to read. Save
them as plain text, and load them as ASCII files.
Other types of binary data files that you might encounter include:
- netcdf: This is a
self-documenting file format that is commonly
used for geophysical data. The files are easy to read into
Matlab if you install the netcdf libraries on your system and the
netcdf toolbox. We'll steer clear of these for this class.
- hdf: This is
another self-documenting file format, though it
comes in different flavors. NASA decided on one version of hdf as
a standard for satellite data collected by the Earth Observation
Satellites. Matlab has an hdf interface that might now work for
NASA hdf files.
- raw binary: These
might include output files from computer simulations, or data files
archived in a compact format. Binary files are often classed as
"big-endian" or "little-endian". The terminology is an allusion
to Jonathan Swift's Gulliver's
Travels, and refers to the arbitrary standards used to write
binary data on different computer systems. In general, PCs use
one format; high end unix workstations often use the other. To
read these files in Matlab, you can do the following:
- fid=fopen('file.bin','r');
or one of the following if the file was written on a different type of
computer: fid=fopen('file.bin','r','l');
or fid=fopen('file.bin','r','b');
- a=fread(fid,'float');
to read the entire file. Check the fread documentation to find
out how to read part of a file, or to read files that have some
floating point numbers and some integers.
- fclose(fid); to
clean up and avoid hogging all the input/output channels.
Clearing data: Sometimes
you want to clear data out of memory. To clear everything and
start afresh use:
clear
To clear one or two variables:
clear
x y
Representing a matrix: Matlab
is really designed to work with matrices, so it very naturally stores
data in two dimensional arrays, and will also use arrays of arbitrary
dimension size. If you read in your data as an ASCII file, you
might end up with a big data array containing latitude in the first
column, longitude in the next, temperature and the third, and so forth
sort of like this:
DATA = [latitudes longitudes salinity
temperature velocity(east/west) velocity(north/south)
velocity(vertical)];
That's perfect for some purposes, but if your original data came from a
two dimensional grid---perhaps they represent sea surface temperature
in the tropical Pacific---then we might want to format the data some
other way. Here's one way to handle it:
SST=reshape(DATA(:,4),nlat,nlon);
will create an nlat by nlon array containing the temperature data from
column 4 of A.
Looking at the data:
To look at the contents of a variable, you can always type its
name. You can also specify specific elements of an array.
For example:
>> T(1:5,1:5)
ans =
27.2218
27.2415 27.2729 27.3757 27.4669
27.4352
27.4828 27.4918 27.5867 27.6677
27.6014
27.6939 27.6883 27.7699 27.8425
27.7381
27.8820 27.8640 27.9348 27.9986
27.8452
28.0494 28.0191 28.0774 28.1343
This is useful for some purposes, but we can't tell much about
our temperature data by looking at it this way.
In Matlab, the first index corresponds to the row, and the second to
the column. So the second row is:
>> T(2,1:5)
ans =
27.4352
27.4828 27.4918 27.5867 27.6677
And the second column is:
>> T(1:5,2)
ans =
27.2415
27.4828
27.6939
27.8820
28.0494
On the otherhand, we can treat a two dimensional array as a vector by
using a single index. Thus:
>> T(1:5)
ans =
27.2218
27.4352 27.6014 27.7381 27.8452
This provides the first 5
elements in column 1, i.e. T(1:5,1).
More on plotting:
The Matlab image functions order arrays like mathematical matrices with
coordinate (1,1) in the upper left corner. Data tend to start
with the smallest latitude and longitude values, which should be mapped
in the lower left corner. To make your Matlab image plot look
correct, you can use axis xy,
as above, or you can flip the matrix top-to-bottom: imagesc(lon_t,lat_t,flipud(T));
controlling details of your plots:
As you work with more data, you may want to control the types of lines
you use for plots, line color, or even the range of colors in an image
plot. Lots of information is available from the help files.
Here are a few pointers.
For simple line plots, lines are ordered
blue-green-red-cyan-magenta-yellow-black, but you can override this by
specifying a single letter color code (e.g. plot(DATA2(:,1),DATA2(:,2),'r') for
a red line.) You can also specify a line type as solid
('-'), dashed ('--'), dotted (':'), or dash-dotted ('-.').
To plot data as points (particularly useful when your data aren't
ordered sequentially), make a scatter plot by specifying a symbol
only:
plot(DATA2(:,1),DATA2(:,2),'o')
to plot circles.
If you start creating a plot and want to keep adding to it, use the hold on command to prevent
Matlab from erasing the first plot when you add a new line and the hold off command to revert to
normal mode. The command clf
will clear everything out of a figure window to let you start again.
colormaps: In Matlab,
the default colormap for contour and image plots is a blue
to red spectrum, but you can override this. To change the
colormap used for contour or image plots, you can specify a different
basic color map by typing, for example, colormap(cool). Other
colormaps include hsv, prism, gray, hot, cool, copper, flag, pink,
bone, and jet (the default).
Sometimes, you want to make sure that NaNs don't end up shaded the same
color as useful data points, so you can force values at the end of your
range to be white or black, for example.
cmap2=[[1 1 1]; colormap; [1 1
1]];
colormap(cmap2);
You might have to fix the limits of your colors to keep real data from
also being whited out. To get black where you had no data, you'd
use [0 0 0].
Plotting vector quantities:
Velocity vectors can be a little subtle. The Matlab function
quiver will plot vector quantities. So:
quiver(lon_t,lat_t,U,V)
plots a map of vector velocities. The results are a little
deceptive. If you change the aspect ratio of the plot, the
vectors appear to change direction, so the on screen vectors aren't
telling you the precise direction of the current.
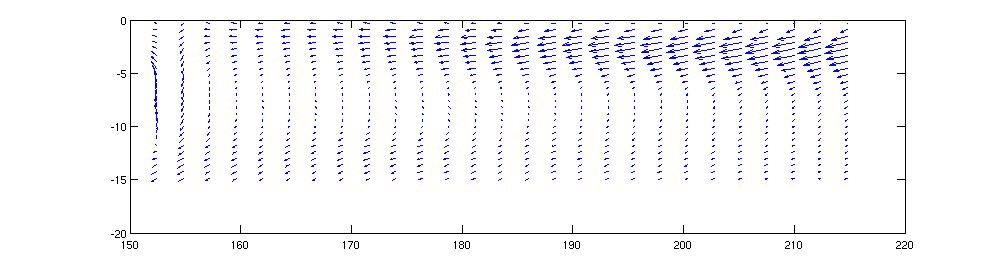
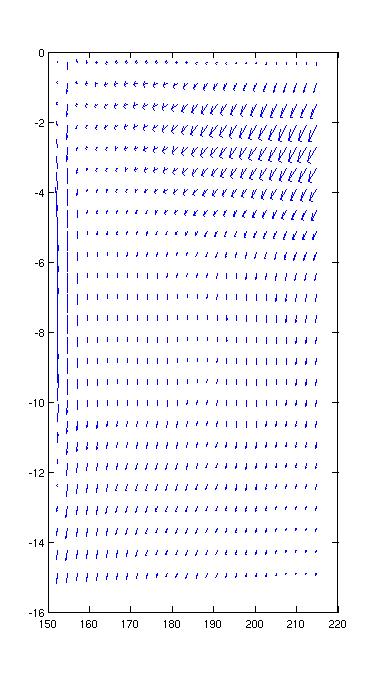
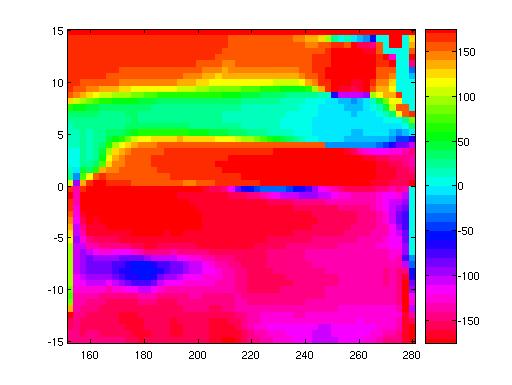
We can also compute the angular direction of the current and plot that
as an image:
theta=atan2(V,U)*180/(pi);
Here we use atan2 rather than atan, because we want our angles
to go from 0 to 360 degrees.
Since 0 and 360 degrees are equivalent, it's good to choose a
colorscale where the colors are 0 and 360 are nearly the same.
The clrscl.m function (written by an SIO
researcher) provides one way to fix a colorscale
appropriately. Compare:
imagesc(lon_t,lat_t,theta);
axis xy; colorbar
with
colormap(clrscl('rmbcgyr',36))
imagesc(lon_t,lat_t,theta); axis
xy; colorbar
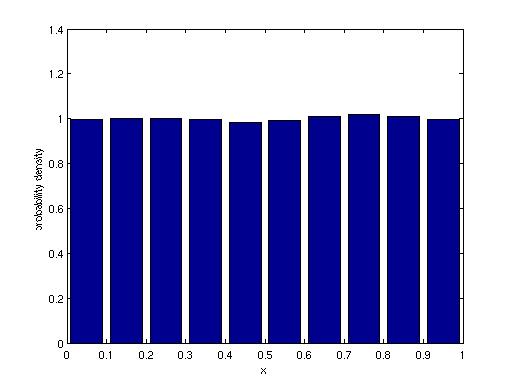
Probability density functions and
histograms: We looked at the hist function last
week. How do you convert a histogram into a probability density
function?
x=rand(20000,1);
[a,b]=hist(x,0.05:.1:.95);
Now we can make a bar plot, typing
bar(b,a)
We can also modify the bar plot to give us what we need for a pdf:
bar(b,a/sum(a)/.1);
% where 0.1 is the interval dx
xlabel('x'); ylabel('probability
density')
You can get rid of the space between the bars by supplying a
width parameter to bar:
bar(b,a/sum(a)/.1,1);
In contrast to this procedure, the Matlab function pdf.m returns the
functional form of a specified probability density function.
Logic: Sometimes we want
to test for a particular condition. In Matlab, conditional
statements can use if, while, or switch. These test whether an
expression is true or false. Expressions test the relationship
between two quantities. The help page for relop explains
everything. Here are some examples.
To test whether T is less than 999, use
(T<999).
To test whether T is equal to 999, use (T==999).
To test whether T is not equal to 999, use (T~=999).
To test whether T and S are both not equal to 999, use (T~=999 &
S~=999).
To test whether T or S is equal to 999, use (T==999 | S == 999).
To test whether T is NaN, use (isnan(T)).
Automating a procedure: Last
week we looked at for loops. Here are some other ways to make a
calculation loop:
run a while loop: if you
prefer to set a logical test to figure out how many times to run your
loop, you can use a while loop. For the example above, you could
say:
i=3;
while i<size(A,2)
array=reshape(A(:,i),nlat,nlon);
figure(i-2)
imagesc(array,lat,lon)
i=i+1;
end
use Matlab's vectorization
capabilities: For many purposes, Matlab will naturally
loop through a whole set of variables without requiring a loop.
For example, if you wanted to compute the mean and standard deviation
of each column in A, you might be tempted to create a loop:
for
i=3:8
m(i-2)=mean(A(:,i));
s(i-2)=std(A(:,i));
end
But instead you could say:
m=mean(A(:,3:8));
s=std(A(:,3:8));
This second option not only requires less code, but it is also
computationally faster.



